| bit | Xplor-NIH | VMD-XPLOR |
|---|
|
| Xplor-NIH home Documentation |
Next: Requirements Up: Distance Restraints Previous: Distance Restraints
Syntax
- NOE {
 noe-statement
noe-statement } END
} END - is invoked from the main level of X-PLOR.
 noe-statement
noe-statement :==
:==-
- ASSIgn
 selection
selection
 selection
selection
 real
real
 real
real
 real
real [OR
[OR  selection
selection
 selection
selection
 ]
adds a distance restraint between the two
selected sets of atoms to the NOE database.
The interpretation of the real numbers depends on the
particular type of restraining function (see Section
20.3). For the POTEntial=BIHA, SQUA, and SOFT
options
the
]
adds a distance restraint between the two
selected sets of atoms to the NOE database.
The interpretation of the real numbers depends on the
particular type of restraining function (see Section
20.3). For the POTEntial=BIHA, SQUA, and SOFT
options
the  real
real numbers are
numbers are 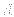 ,
, 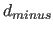 , and
, and 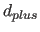 (see also Section 20.4) (default: none). One or more
optional OR clauses can be appended to the the ASSIgn statement,
each introducing an additional ambiguous pair of selections to the
given assignment. See Section 20.2.1 for an explanation
of how these additional selections are used.
(see also Section 20.4) (default: none). One or more
optional OR clauses can be appended to the the ASSIgn statement,
each introducing an additional ambiguous pair of selections to the
given assignment. See Section 20.2.1 for an explanation
of how these additional selections are used.
- ASYMptote
 *class*
*class*
 real
real is the slope
is the slope 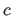 of the
asymptote of the soft-square function
(Eq. 20.12) (default: 0).
of the
asymptote of the soft-square function
(Eq. 20.12) (default: 0).
- AVERaging
 *class*
*class* R-6
R-6 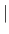 R-3
R-3 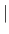 SUM
SUM 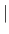 CENTer
specifies the averaging method (Section 20.2)
(default: R-6).
CENTer
specifies the averaging method (Section 20.2)
(default: R-6).
- BHIG
 *class*
*class*
 real
real
- specifies the height of the potential barriers in the high dimensional restraining function (Eq. 20.17), only for POTEntial=HIGH (default: 0.01).
- CEILing=
 real
real
- specifies the ceiling value for energy constants; it applies to most restraining functions (Section 20.3) (default: 30).
- CLASsification
 class-name
class-name
- partitions the distance restraints database; it applies to all subsequent ASSIgn entries until another CLASS entry is issued (default: NIL).
- COUNtviol
 class
class
- counts violations in class and stores the smallest violation in the symbol “$RESULT"; action is taken when this statement is issued.
- DISTribute
 class1
class1
 class2
class2
 real
real
- distributes
distance restraints between two classes: the
restraint will be put in
 class1
class1 if the distance
is less than
if the distance
is less than 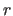 (the specified real number);
it will be put in
(the specified real number);
it will be put in  class2
class2 if the
distance is greater than or equal to
if the
distance is greater than or equal to 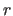 .
Action is taken only when this statement is issued.
.
Action is taken only when this statement is issued.
- MONOmers
 *class*
*class*
 integer
integer specifies the
number of monomers for the AVERage=SUM option
(Section 20.2) (default: 1).
specifies the
number of monomers for the AVERage=SUM option
(Section 20.2) (default: 1).
- NCOUnt
 *class*
*class*
 integer
integer specifies
number of assign statements for the high dimensional restraining
function (Eq. 20.17); only even values are
allowed (default: 2).
specifies
number of assign statements for the high dimensional restraining
function (Eq. 20.17); only even values are
allowed (default: 2).
- NREStraints=
 integer
integer
- is a required parameter that specifies the maximum expected number of distance restraints. It has to be greater than or equal to the actual number of restraints. The parameter is used to allocate space dynamically for the restraint list (default: 200).
- POTEntial
 *class*
*class* BIHArmonic
BIHArmonic 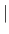 LOGNormal
LOGNormal 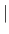 SQUAre
SQUAre  SOFTsquare
SOFTsquare  SYMMetry
SYMMetry
 HIGH
HIGH  3DPO
specifies the restraining function (Section 20.3)
(default: BIHArmonic).
3DPO
specifies the restraining function (Section 20.3)
(default: BIHArmonic).
- PREDict
- {
 predict-statement
predict-statement } END can be used to
predict interproton distances based on the Cartesian coordinates in the
MAIN coordinate set. It
lists all distances between the FROM and the TO selections of
atoms that are within the range CUTOn ...CUTOff. Distances
are
} END can be used to
predict interproton distances based on the Cartesian coordinates in the
MAIN coordinate set. It
lists all distances between the FROM and the TO selections of
atoms that are within the range CUTOn ...CUTOff. Distances
are 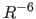 averaged over the GROUps that are
defined in the topology file for each residue
(see Section 3.1.1).
Current distance restraint information is overwritten by this statement.
averaged over the GROUps that are
defined in the topology file for each residue
(see Section 3.1.1).
Current distance restraint information is overwritten by this statement.
- PRINt THREshold=
 real
real
- prints distance violations
greater than the specified real number. This statement also
declares two symbols, $RESULT and $VIOLATIONS, which correspond to the
rms deviation of the distances and the number of violations greater than
the THREshold value, respectively.
The definition of the deviation depends on which restraining function
is specified: in the case of the SQUAre-well and SOFTsquare
functions, the deviation refers to the difference
between the actual distance and the upper or lower
bound if the distance is outside the error range; in
the case of the BIHArmonic function,
the deviation refers to the difference between the actual
distance and the target distance.
(A more precise definition is
given by
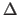 in Section 20.3.)
in Section 20.3.)
- RESEt
- erases the current NOE database.
- RSWItch
 *class*
*class*
 real
real
- is the switch parameter
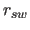 between
the square-well and the asymptote (Eq. 20.12)
(default: 10).
between
the square-well and the asymptote (Eq. 20.12)
(default: 10).
- SCALe
 *class*
*class*
 real
real
- scales the restraint class (Section 20.3) (default: 1).
- SOEXponent
 *class*
*class*
 real
real
- is an exponent; it is for the soft-square function (Eq. 20.12) (default: 2).
- SQCOnstant
 *class*
*class*
 real
real
- is an additional scale factor; it is for the square-well or soft-square function (Eqs. 20.10 and 20.12) (default: 20).
- SQEXponent
 *class*
*class*
 real
real
- is an exponent; it is for the square-well or soft-square function (Eqs. 20.10 and 20.12) (default: 2).
- SQOFfset
 *class*
*class*
 real
real
- applies
a negative offset value to all upper bounds
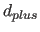 of the
specified class(es). This offset is
applied to the square-well or soft-square functions
(Eqs. 20.10 and 20.12) and to the
bounds used by distance geometry (see Chapter 37) (default: 0).
of the
specified class(es). This offset is
applied to the square-well or soft-square functions
(Eqs. 20.10 and 20.12) and to the
bounds used by distance geometry (see Chapter 37) (default: 0).
- TEMPerature=
 real
real
- is the temperature in K; it is only for the biharmonic function (Eq. 20.7) (default: 300).
 predict-statement
predict-statement :==
:==-
- CUTOff=
 real
real
- specifies an upper limit for acceptable interproton distances (default: 10.0 Å).
- CUTOn=
 real
real
- specifies a lower limit for acceptable interproton distances (default: 0.0 Å).
- FROM=
 selection
selection
- specifies an atom selection (default: none).
- TO=
 selection
selection
- specifies an atom selection (default: none).
- CUTOff=
Subsections Xplor-NIH 2025-11-07
