| bit | Xplor-NIH | VMD-XPLOR |
|---|
|
| Xplor-NIH home Documentation |
Next: Requirements Up: Metric Matrix Distance Geometry Previous: Metric Matrix Distance Geometry
Syntax
- MMDG {
 mmdg-statement
mmdg-statement } END
} END - is invoked from the main level of X-PLOR.
 mmdg-statement
mmdg-statement :==
:==-
- BACCuracy=
 real
real
- specifies the value (in Å) that is added to and subtracted from each bond length parameter to give upper and lower bounds on the 1-2 distance (default: 0.01 Å).
- EXPOnent
 integer
integer
- specifies the exponent for the DG restraint term for regularization (see Section 37.4).
- GROUp=
 selection
selection
 real
real
- selects the atoms that form a rigid group, based on the main coordinate set (see Section 6.1), and the value (in Å) that is added to and subtracted from each interatomic distance calculated from that rigid group. Note that the GROUp statement uses the main coordinate set, in contrast to the REFErence=COORdinate statement and the pseudoatom correction, which use the reference coordinate set. Multiple specifications of GROUps define multiple rigid groups. The group selections need not be disjoint. By default, no rigid groups are defined. The maximum number of groups is currently limited to NATOM, the number of atoms in the molecular structures; the maximum number of all selected atoms in all GROUp statements combined is currently limited to 10*NATOM.
- IACCuracy=
 real
real
- specifies the value (in degrees)
that is added to and subtracted from each improper angle parameter
defined by atoms i,j,k,l
to give upper and lower bounds on the distance
between atoms i and l (default: 2
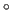 ).
).
- METRization
 selection
selection
 integer
integer
- performs metrization. If this statement is not specified, only bound-smoothing is performed. The statement selects the atoms from which distances can be chosen during the retightening phase of the partial metrization protocol. The integer tells exactly how many of the selected atoms will be used in the retightening phase, which is useful for partial metrization (see Kuszewski, Nilges, and Brünger, 1992). If the integer is greater than the number of selected atoms, it is automatically reduced to that number. All other distances are chosen after these and without retightening (although bound smoothing is performed for all atoms).
- ORDEred
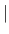 RANDom
RANDom - are exclusive flags that set the metrization order to either ordered (by internal atom number) or random (default: random).
- PACCuracy=
 real
real
- specifies the value (in degrees)
that is to be added to and subtracted from each dihedral angle
parameter defined by atoms i,j,k,l
to give upper and lower bounds on the distance between atoms i and l
(default: 2
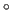 ).
).
- READBOunds=
 filename
filename
- reads smoothed-bound matrices and then performs metrization and embedding.
- RECALLbounds
- recalls smoothed-bound matrices from memory and then performs metrization and embedding.
- REFErence=PARAmeter
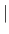 COORdinates
COORdinates - uses the parameter database to generate distances for covalent bonds, bond angles, improper angles, and dihedral angles with a single minimum if PARAmeter is specified. If COORdinates is specified, the coordinates from the reference coordinate set (see Section 6.1) are used to obtain the distances. In this case, the accuracy of the distance bounds is specified by BACCuracy, regardless of whether a bond and bond angle define the particular distance. Dihedral angles with multiple minima, nonbonded parameters, and distance and dihedral angle restraints are unaffected by this option (default: PARAmeter).
- SCALe
 real
real
- specifies the scale factor for the DG restraint term for regularization (see Section 37.4).
- SELEction
 selection
selection
- specifies the atoms that are used in the DG restraint for regularization (see Section 37.4) (default: (NOT ALL) ).
- SHORtest-path-algorithm=AUTO
 FULL
FULL  SPARse
SPARse - specifies
the algorithm for the shortest path used during metrization and bound smoothing. When AUTO is specified, the program automatically tries to decide which algorithm is more appropriate. If FULL is specified, the Dijkstra algorithm is used. If SPARse is specified, a modified version of the Dijkstra algorithm operates on the (sparse) tree of known distances (default: AUTO). - STOREBounds
- stores smoothed-bound matrices in memory, and no metrization or embedding is performed. This option saves CPU time when multiple metrizations and embeddings are carried out using the same distance constraints. The option is also required when using the distance geometry restraint term (Section 37.4), which allows the user to regularize coordinates using distance bounds information only.
- SUBStructure
 selection
selection
- specifies the atoms to be embedded (default: (ALL) ).
- TACCuracy=
 real
real
- specifies the value (in degrees) that is
to be added to and subtracted from each bond angle parameter to give
upper and lower bounds on the 1-3 distance (default: 2
 ).
).
- VERBose=
 logical
logical
- switches verbose mode on and off. If verbose mode is on, many status reports are printed (default: off).
- WRITEBounds=
 filename
filename
- writes smoothed-bound matrices to the specified file; no metrization or embedding is performed.
- BACCuracy=
Upon execution of the END statement, the program performs all steps of distance geometry (initialization, metrization, smoothing, and embedding), unless STOREBounds, WRITEBounds, RECALLbounds, or READBOunds is specified.
Xplor-NIH 2025-11-07
