| bit | Xplor-NIH | VMD-XPLOR |
|---|
|
| Xplor-NIH home Documentation |
Next: Requirements Up: Setup of the Relaxation Previous: Setup of the Relaxation
Syntax
- RELAxation {
 relaxation-statement
relaxation-statement } END
} END - is
invoked from the
main level of X-PLOR.  relaxation-statement
relaxation-statement :==
:==
- ASSIgn
 selection
selection
 selection
selection {
{
![${<\mbox{real}>[<\mbox{real}>[<\mbox{real}>]]}$](img790.png) }
adds a single cross-peak intensity or a
buildup curve between the two
specified selected sets of spins. The number of mixing times
read depends on the number of CLASs statements issued before the
ASSIgn statement.
Error estimates can be specified. The
format depends on the ERROr_input statement. For
examples, see Section 39.7.
}
adds a single cross-peak intensity or a
buildup curve between the two
specified selected sets of spins. The number of mixing times
read depends on the number of CLASs statements issued before the
ASSIgn statement.
Error estimates can be specified. The
format depends on the ERROr_input statement. For
examples, see Section 39.7.
- AVERage
 real
real
- specifies an exponent for calculating the average distance to an UNREsolved group of protons (default: 6).
- CALIbrate {
 calibrate-statement
calibrate-statement } END
} END - calculates the calibration factor between the observed and calculated intensities. Prior to the calibration, this command computes the NOE intensities from the current atomic model unless VALUe is specified.
 calibrate-statement
calibrate-statement :==
:==-
- AUTOmatic=
 ON,OFF
ON,OFF
- sets the calibration factor, if ON, every step during the refinement. At present, this allows the calculation of the gradient only approximately. Therefore, it should be turned OFF during minimization.
- GROUp=
 CLASs,GROUp,ALL
CLASs,GROUp,ALL
- for CLASS, calculates the calibration factor separately for each classification; for ALL, one overall calibration factor is used (default: CLASs).
- QUALity=
 real
real
- specifies that cross peaks for which the relative error estimate exceeds the specified value are not used (default: 1).
- REFErence=
 ALL,REFErence
ALL,REFErence
- for ALL, uses all peaks that meet the criterion set with the EXCLude option for the calculation of the calibration factor; for REFErence, uses only those that were flagged as reference peaks during input (default: ALL).
- VALUe class
 *real*
*real*
- sets a value for the calibration factor. Note: if the calibration factor is set by VALUe for a class, it is not recalculated for other classes in the same CALIbrate statement.
- AUTOmatic=
- CLASsification
 class-name
class-name
- partitions an intensity table; it applies to all following ASSIgn entries until another CLASS entry is issued. In contrast to the NOE CLASs statement, the RELAxation CLASs statement can be used to switch between classes during the input of cross-peak intensities. Several class statements can be issued without an intervening ASSIgn statement. This allows the input of buildup curves (default: NIL).
- CLGR (CLass_GRoup)
 *class*
*class*
 group
group
- enters a CLASs into a GROUp of spectra (default: none).
- CUTOff
- {
 cutoff-statement
cutoff-statement } END applies a cutoff to
relaxation matrix and/or gradient calculation. When a cutoff is in
effect, for each peak
on the data list a separate small relaxation matrix is set up,
including only spins
within a range of the spin pair specified by the cutoff.
The spectrum and the gradient are calculated with this
small relaxation matrix.
} END applies a cutoff to
relaxation matrix and/or gradient calculation. When a cutoff is in
effect, for each peak
on the data list a separate small relaxation matrix is set up,
including only spins
within a range of the spin pair specified by the cutoff.
The spectrum and the gradient are calculated with this
small relaxation matrix.
 cutoff-statement
cutoff-statement :==
:== -
- MODE=
 NONE,GRADient,ALL
NONE,GRADient,ALL
- applies no cutoff, cutoff only for the gradient calculation, or cutoff for both intensity and gradient calculation (default: NONE).
- RELAtive_value=
 real
real
- specifies a cutoff relative to the actual distance between the two spins in Å (default: 0).
- VALUe=
 real
real
- specifies the actual value of the cutoff, in Å (default: 6).
- MODE=
- EEXPonent
 real
real
- is an exponent
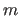 of the WELL or
PARAbolic energy function (default: 2).
of the WELL or
PARAbolic energy function (default: 2).
- ERROr_input
- {
 error-input-statement
error-input-statement } END
defines an error input format for the following ASSIgn statement.
It remains in effect until another ERROr_input command is
issued.
} END
defines an error input format for the following ASSIgn statement.
It remains in effect until another ERROr_input command is
issued.
 errorinput-statement
errorinput-statement :==
:==-
- INPUt=
 UNIForm,RANGe,PLUSminus
UNIForm,RANGe,PLUSminus
- specifies the input format (0, 1, or 2 error estimates) for the following ASSIgn statements (default: RANGe): for UNIForm, no error estimate is expected by the program, for RANGe, one error estimate, and for PLUSminus, two, where the first is an error estimate on the minus side, the second on the plus side.
- MODE=
 ABSOlute,RELAtive
ABSOlute,RELAtive
- interprets the error estimate in the ASSIgn statements as absolute or relative (default: ABSOlute).
- VALUe=
 real
real
- enters the actual value for UNIForm option (default: 0).
- INPUt=
- GROUp
 group-name
group-name
- defines a group of several
CLASses that are recorded with the same physical conditions,
like solvent(H
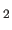 O, D
O, D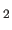 O), field strength, and temperature
(default: NIL).
O), field strength, and temperature
(default: NIL).
- IEXPonent
 real
real
- specifies an exponent
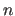 of the calculated and
observed intensities in the energy function (default: 1).
of the calculated and
observed intensities in the energy function (default: 1).
- IWEIght
- {
 iweight-statement
iweight-statement } END applies
individual weights
} END applies
individual weights 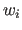 to each single term in the energy function.
At the moment, the only supported options are weights equal to the inverse
of
powers of the observed intensity (i.e.,
to each single term in the energy function.
At the moment, the only supported options are weights equal to the inverse
of
powers of the observed intensity (i.e.,
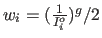 ).
If the exponent
).
If the exponent 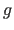 matches the exponent
matches the exponent 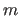 of the energy
function, the weighting is equivalent to refining relative intensity
differences. The minimum weight applied is always 1.
of the energy
function, the weighting is equivalent to refining relative intensity
differences. The minimum weight applied is always 1.
 iweight-statement
iweight-statement :==
:== -
- CUTOff=
 real
real
- is the maximum weight applied (default: 10000).
- EXPOnent=
 real
real
- specifies an exponent
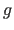 of the
observed intensity in the weight function
of the
observed intensity in the weight function
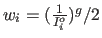 (default: 2).
(default: 2).
- QUALity=
 real
real
- stipulates that cross peaks for which the relative error estimate exceeds the specified value are not used in determining the maximum weight (default: 1).
- CUTOff=
- MINIntensity
 *class*
*class*
 real
real
- sets the minimum detected intensity for each class. The specified value is used as the “zero" for the intensities (default: smallest number used by X-PLOR).
- NREStraints
 integer
integer
- is the required parameter that specifies the maximum expected number of intensities. It has to be greater than or equal to the actual number. The parameter is used to assign space dynamically for the intensity list (default: 200).
- OCCUpancy
 group
group
 selection
selection
 real
real
- sets the occupancy for selected spins (for example, 0.9 for exchanging amid protons) (default: 1.0).
- OMEGa
 *group*
*group*
 real
real
- specifies the spectrometer frequency, in Hz
(default: 500*10
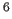 ).
).
- OMIT
 group
group
 selection
selection
- is an obsolete statement that was used in a prerelease version of the relaxation matrix code of X-PLOR. It will be removed in a future version. The statement deletes spins from the relaxation matrix. This is almost the reverse of the SELEct command, but it leaves any existing base of unresolved groups intact (default: none).
- POTEntial
 WELL,PARAbolic
WELL,PARAbolic
- provides two alternatives: WELL uses a flat-bottom potential with bounds at error estimates, PARAbolic a (generalized) parabola with exponent EEXP and center at the specified intensity (default: WELL).
- PREDict_intensities
 class
class {
{  predict-intensities-statement
predict-intensities-statement } END
calculates a spectrum from coordinates with the parameters valid for the
specified class. Note that this destroys any existing experimental intensity database.
The spectrum can be calculated for parts of a molecular structure.
} END
calculates a spectrum from coordinates with the parameters valid for the
specified class. Note that this destroys any existing experimental intensity database.
The spectrum can be calculated for parts of a molecular structure.
 PREDict_intensities-statement
PREDict_intensities-statement :==
:== -
- CUTOFf
 real
real
- specifies a distance cutoff for cross-peak calculation (default: 10).
- CUTON
 real
real
- specifies a distance cuton for cross-peak calculation (default: 0).
- DIAGonal
 ON,OFF
ON,OFF
- prints diagonal peaks (default: OFF).
- FORMat
 NORMal,LIST
NORMal,LIST
- is an output format; LIST produces an intensity INPUT file (default: NORMal).
- FROM
 selection
selection
- selects the first part of a molecular structure (default: all NMR-active atoms).
- THREshold
 real
real
- specifies the minimum intensity that is printed (default: 0).
- TO
 selection
selection
- selects the second part of a molecular structure (default: all NMR-active atoms).
- CUTOFf
- PRINt_deviations
- {
 print-deviation-statement
print-deviation-statement } END
prints the deviations from observed intensities and calculates
an
} END
prints the deviations from observed intensities and calculates
an 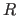 value. It uses the form of the energy function
(WELL or PARAbolic and the value of IEXP) specified
before issuing the statement.
value. It uses the form of the energy function
(WELL or PARAbolic and the value of IEXP) specified
before issuing the statement.
 print-deviation-statement
print-deviation-statement :==
:== -
- FORMat
 NORMal,LIST
NORMal,LIST
- is an output format; LIST produces a listing in the format of an intensity input file.
- SELEct
 selection
selection
- calculates the
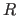 value with only a
subset of the atoms. This allows the calculation of
local
value with only a
subset of the atoms. This allows the calculation of
local 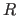 values.
values.
- THREshold
 real
real
- specifies the minimum deviation that is printed, in the arbitrary units of the input intensities. For a complete list of deviations (including 0), THREshold should be set to a negative number (default: 0).
- FORMat
- REFErence
- flags intensities read by the following ASSIgn statements as reference peaks for the calculation of the calibration factor (see CALIbrate statement), and remains in effect until another CLASs statement is issued. This flag allows one to use a subset of peaks for the calibration, e.g., peaks that do not depend very much on the conformation of the molecule. The reference peaks are written into a separate file, and the reference flag is set before reading that file.
- RESEt
- erases the current relaxation database.
- SELEct
 group
group
 selection
selection
- selects the “NMR-active" atoms (default: (HYDRogen) ). At present, only protons can be used in the refinement. The NMR parameters for protons are applied to all selected atoms. This command initializes the array that stores information about unresolved groups (see below). If the command is not issued at the beginning of the relaxation matrix setup, all hydrogens are used by default. Since a hydrogen is defined by its mass in X-PLOR, care should be taken in this case to modify the masses of hydrogens for simulated annealing only after the relaxation matrix parameters have been set up.
- SORT_intensities
- sorts the intensity list so that peaks with different mixing times belonging to the same spin pair are consecutive on the list. If a cutoff is used, a new relaxation matrix generally has to be set up and diagonalized for each peak on the data list. However, if the only difference between two entries on the data list is the mixing time, the relaxation matrix does not have to be recalculated and diagonalized. Sorting can thus save a considerable amount of CPU time.
- TAUCorrelation
 group
group {
{  TAUC-statement
TAUC-statement } END
specifies the correlation time and the order parameter, depending
on the chosen model.
} END
specifies the correlation time and the order parameter, depending
on the chosen model.
 TAUCorrelation-statement
TAUCorrelation-statement :==
:== -
- MODEl
 RIGId
RIGId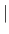 LIPAri
LIPAri
- provides two alternatives: RIGId expects no order parameter; LIPAri expects one overall correlation time and one order parameter. This applies to both the ISOTropic and the VECTor case.
- ISOTropic
![$<\mbox{real}>[<\mbox{real}>]$](img795.png)
- specifies an isotropic correlation time and an order parameter (default: 10 ns, 1).
- RESEt
- initializes a table.
- VECTor
 selection
selection
 selection
selection
![$<\mbox{real}>[<\mbox{real}>]$](img795.png)
- specifies a
correlation time (in seconds) and an order parameter for a specified vector.
- MODEl
- TAUMix
 *class*
*class*
 real
real
- specifies the mixing time
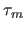 , in seconds (default: 0.1 sec).
, in seconds (default: 0.1 sec).
- TOLErance
 real
real
- if not equal to zero,
approximates the exact calculation of the
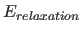 energy function and its first
derivatives with respect to the atomic coordinates
by freezing
energy function and its first
derivatives with respect to the atomic coordinates
by freezing
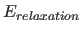 until atoms
have moved by less than TOLErance Å from the point at which
until atoms
have moved by less than TOLErance Å from the point at which
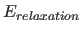 was
last computed. This is particularly useful for
molecular dynamics calculations. It should be set to 0
during minimization to avoid inconsistencies
in the force field (default: 0).
was
last computed. This is particularly useful for
molecular dynamics calculations. It should be set to 0
during minimization to avoid inconsistencies
in the force field (default: 0).
- UPDate
- computes the NOE intensities from the current atomic model.
- UNREsolved
 group
group {
{  unresolved-statement
unresolved-statement } END
replaces protons with unresolved chemical shifts by a single
spin. The distance to an unresolved group is calculated
as
} END
replaces protons with unresolved chemical shifts by a single
spin. The distance to an unresolved group is calculated
as
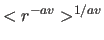 , where the exponent
, where the exponent 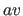 is set with the
AVERage command. A diagonal leakage rate is added
to the relaxation matrix for each unresolved group.
The main use of this command is for methyl groups.
is set with the
AVERage command. A diagonal leakage rate is added
to the relaxation matrix for each unresolved group.
The main use of this command is for methyl groups.
 unresolved-statement
unresolved-statement :==
:== -
- 1-2
- treats each methylene group as an unresolved group of protons. A methylene group is recognized by X-PLOR as two hydrogens bound to a heavy atom.
- METHyl
- treats each methyl group as an unresolved group of protons. A methyl group is recognized by X-PLOR as three hydrogens bound to a heavy atom.
- NONE
- deletes all previously defined unresolved groups.
- SELECT
 selection
selection
- treats the selected atoms as an unresolved group of protons.
- WEIGht
 *class*
*class*
 real
real
- specifies the
energy constant
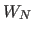 (overall weight)
(default: 1).
(overall weight)
(default: 1).
- ZLEAkage_rate
 *group*
*group*
- specifies the overall diagonal
leakage rate
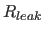 , in sec
, in sec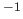 (default: 0).
(default: 0).
