| bit | Xplor-NIH | VMD-XPLOR |
|---|
|
| Xplor-NIH home Documentation |
Next: Distance Matrix Analysis Up: Cartesian Coordinates Previous: Example
Rms Differences between Coordinates
The following example shows how to compute all rms
differences (rmsd) between two coordinate sets (one in the main
set, one in the comparison set). The rmsds are rms-averaged over
all atoms of each residue. The user is also referred to
Sections 38.8 and 38.9
for computation of averages and
rmsds of a family of coordinate sets.
X-PLOR produces the file “rmsdplot.list", which can be interpreted by Mathematica:
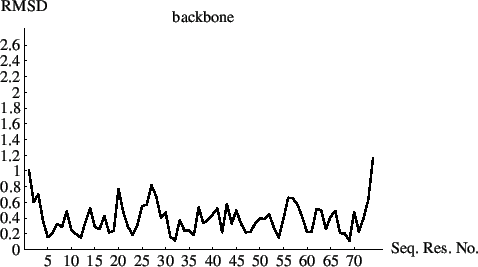
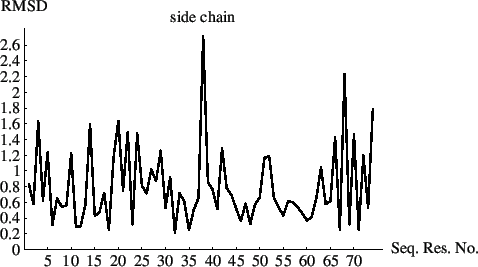
The resulting plots of the rmsds as a function of sequential residue number are shown in Figs. 6.1 and 6.2.
The correspondence between sequential residue number and actual residue identifier can be established by using vector statements (see Section 5.4).
Xplor-NIH 2025-11-07